Processing Data_Michelle_RNAseq_2025
gene_x 0 like s 9 view s
Tags: processing, pipeline
-
Targets
The experiment we did so far: I have two strains: 1. 1457 wildtype 2. 1457Δsbp (sbp knock out strain) I have grown these two strains in two media for 2h (early biofilm phase, primary attachment), 4h (biofilm accumulation phase), 18h (mature biofilm phase) respectively 1. medium TSB -> nutrient-rich medium: differences in biofilm formation and growth visible (sbp knockout shows less biofilm formation and a growth deficit) 2. medium MH -> nutrient-poor medium: differences between wild type more obvious (sbp knockout shows stronger growth deficit) Our idea/hypothesis of what we hope to achieve with the RNA-Seq: Since we already see differences in growth and biofilm formation and also differences in the proteome (through cooperation with mass spectrometry), we also expect differences in the transcription of the genes in the RNA-Seq. Could you analyze the RNA-Seq data for me and compare the strains at the different time points? But maybe also compare the different time points of one strain with each other? The following would be interesting for me: - PCA plot (sample comparison) - Heatmaps (wild type vs. sbp knockout) - Volcano plots (significant genes) - Gene Ontology (GO) analyses -
Download the raw data
aws s3 cp s3://xxxx-xxxxxxx/ ./ --recursive -
Prepare raw data
mkdir raw_data; cd raw_data #Δsbp->deltasbp #1457.1_2h_MH,WT,MH,2h,1 ln -s ../F25A430000603-01_STAkimxP/1457.1_2h_MH/1457.1_2h_MH_1.fq.gz WT_MH_2h_1_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457.1_2h_MH/1457.1_2h_MH_2.fq.gz WT_MH_2h_1_R2.fastq.gz #1457.2_2h_ ln -s ../F25A430000603-01_STAkimxP/1457.2_2h_MH/1457.2_2h_MH_1.fq.gz WT_MH_2h_2_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457.2_2h_MH/1457.2_2h_MH_2.fq.gz WT_MH_2h_2_R2.fastq.gz #1457.3_2h_ ln -s ../F25A430000603-01_STAkimxP/1457.3_2h_MH/1457.3_2h_MH_1.fq.gz WT_MH_2h_3_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457.3_2h_MH/1457.3_2h_MH_2.fq.gz WT_MH_2h_3_R2.fastq.gz #1457.1_4h_ ln -s ../F25A430000603-01_STAkimxP/1457.1_4h_MH/1457.1_4h_MH_1.fq.gz WT_MH_4h_1_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457.1_4h_MH/1457.1_4h_MH_2.fq.gz WT_MH_4h_1_R2.fastq.gz #1457.2_4h_ ln -s ../F25A430000603-01_STAkimxP/1457.2_4h_MH/1457.2_4h_MH_1.fq.gz WT_MH_4h_2_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457.2_4h_MH/1457.2_4h_MH_2.fq.gz WT_MH_4h_2_R2.fastq.gz #1457.3_4h_ ln -s ../F25A430000603-01_STAkimxP/1457.3_4h_MH/1457.3_4h_MH_1.fq.gz WT_MH_4h_3_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457.3_4h_MH/1457.3_4h_MH_2.fq.gz WT_MH_4h_3_R2.fastq.gz #1457.1_18h_ ln -s ../F25A430000603-01_STAkimxP/1457.1_18h_MH/1457.1_18h_MH_1.fq.gz WT_MH_18h_1_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457.1_18h_MH/1457.1_18h_MH_2.fq.gz WT_MH_18h_1_R2.fastq.gz #1457.2_18h_ ln -s ../F25A430000603-01_STAkimxP/1457.2_18h_MH/1457.2_18h_MH_1.fq.gz WT_MH_18h_2_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457.2_18h_MH/1457.2_18h_MH_2.fq.gz WT_MH_18h_2_R2.fastq.gz #1457.3_18h_ ln -s ../F25A430000603-01_STAkimxP/1457.3_18h_MH/1457.3_18h_MH_1.fq.gz WT_MH_18h_3_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457.3_18h_MH/1457.3_18h_MH_2.fq.gz WT_MH_18h_3_R2.fastq.gz #1457dsbp1_2h_ ln -s ../F25A430000603-01_STAkimxP/1457dsbp1_2h_MH/1457dsbp1_2h_MH_1.fq.gz deltasbp_MH_2h_1_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457dsbp1_2h_MH/1457dsbp1_2h_MH_2.fq.gz deltasbp_MH_2h_1_R2.fastq.gz #1457dsbp2_2h_ ln -s ../F25A430000603-01_STAkimxP/1457dsbp2_2h_MH/1457dsbp2_2h_MH_1.fq.gz deltasbp_MH_2h_2_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457dsbp2_2h_MH/1457dsbp2_2h_MH_2.fq.gz deltasbp_MH_2h_2_R2.fastq.gz #1457dsbp3_2h_ ln -s ../F25A430000603-01_STAkimxP/1457dsbp3_2h_MH/1457dsbp3_2h_MH_1.fq.gz deltasbp_MH_2h_3_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457dsbp3_2h_MH/1457dsbp3_2h_MH_2.fq.gz deltasbp_MH_2h_3_R2.fastq.gz #1457dsbp1_4h_ ln -s ../F25A430000603-01_STAkimxP/1457dsbp1_4h_MH/1457dsbp1_4h_MH_1.fq.gz deltasbp_MH_4h_1_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457dsbp1_4h_MH/1457dsbp1_4h_MH_2.fq.gz deltasbp_MH_4h_1_R2.fastq.gz #1457dsbp2_4h_ ln -s ../F25A430000603-01_STAkimxP/1457dsbp2_4h_MH/1457dsbp2_4h_MH_1.fq.gz deltasbp_MH_4h_2_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457dsbp2_4h_MH/1457dsbp2_4h_MH_2.fq.gz deltasbp_MH_4h_2_R2.fastq.gz #1457dsbp3_4h_ ln -s ../F25A430000603-01_STAkimxP/1457dsbp3_4h_MH/1457dsbp3_4h_MH_1.fq.gz deltasbp_MH_4h_3_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457dsbp3_4h_MH/1457dsbp3_4h_MH_2.fq.gz deltasbp_MH_4h_3_R2.fastq.gz #1457dsbp118h_ ln -s ../F25A430000603-01_STAkimxP/1457dsbp118h_MH/1457dsbp118h_MH_1.fq.gz deltasbp_MH_18h_1_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457dsbp118h_MH/1457dsbp118h_MH_2.fq.gz deltasbp_MH_18h_1_R2.fastq.gz #1457dsbp218h_ ln -s ../F25A430000603-01_STAkimxP/1457dsbp218h_MH/1457dsbp218h_MH_1.fq.gz deltasbp_MH_18h_2_R1.fastq.gz ln -s ../F25A430000603-01_STAkimxP/1457dsbp218h_MH/1457dsbp218h_MH_2.fq.gz deltasbp_MH_18h_2_R2.fastq.gz #1457.1_2h_ ln -s ../F25A430000603_STAmsvaP/1457.1_2h_TSB/1457.1_2h_TSB_1.fq.gz WT_TSB_2h_1_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457.1_2h_TSB/1457.1_2h_TSB_2.fq.gz WT_TSB_2h_1_R2.fastq.gz #1457.2_2h_ ln -s ../F25A430000603_STAmsvaP/1457.2_2h_TSB/1457.2_2h_TSB_1.fq.gz WT_TSB_2h_2_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457.2_2h_TSB/1457.2_2h_TSB_2.fq.gz WT_TSB_2h_2_R2.fastq.gz #1457.3_2h_ ln -s ../F25A430000603_STAmsvaP/1457.3_2h_TSB/1457.3_2h_TSB_1.fq.gz WT_TSB_2h_3_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457.3_2h_TSB/1457.3_2h_TSB_2.fq.gz WT_TSB_2h_3_R2.fastq.gz #1457.1_4h_ ln -s ../F25A430000603_STAmsvaP/1457.1_4h_TSB/1457.1_4h_TSB_1.fq.gz WT_TSB_4h_1_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457.1_4h_TSB/1457.1_4h_TSB_2.fq.gz WT_TSB_4h_1_R2.fastq.gz #1457.2_4h_ ln -s ../F25A430000603_STAmsvaP/1457.2_4h_TSB/1457.2_4h_TSB_1.fq.gz WT_TSB_4h_2_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457.2_4h_TSB/1457.2_4h_TSB_2.fq.gz WT_TSB_4h_2_R2.fastq.gz #1457.3_4h_ ln -s ../F25A430000603_STAmsvaP/1457.3_4h_TSB/1457.3_4h_TSB_1.fq.gz WT_TSB_4h_3_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457.3_4h_TSB/1457.3_4h_TSB_2.fq.gz WT_TSB_4h_3_R2.fastq.gz #1457.1_18h_ ln -s ../F25A430000603_STAmsvaP/1457.1_18h_TSB/1457.1_18h_TSB_1.fq.gz WT_TSB_18h_1_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457.1_18h_TSB/1457.1_18h_TSB_2.fq.gz WT_TSB_18h_1_R2.fastq.gz #1457.2_18h_ ln -s ../F25A430000603_STAmsvaP/1457.2_18h_TSB/1457.2_18h_TSB_1.fq.gz WT_TSB_18h_2_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457.2_18h_TSB/1457.2_18h_TSB_2.fq.gz WT_TSB_18h_2_R2.fastq.gz #1457.3_18h_ ln -s ../F25A430000603_STAmsvaP/1457.3_18h_TSB/1457.3_18h_TSB_1.fq.gz WT_TSB_18h_3_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457.3_18h_TSB/1457.3_18h_TSB_2.fq.gz WT_TSB_18h_3_R2.fastq.gz #1457dsbp1_2h_ ln -s ../F25A430000603_STAmsvaP/1457dsbp1_2hTSB/1457dsbp1_2hTSB_1.fq.gz deltasbp_TSB_2h_1_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457dsbp1_2hTSB/1457dsbp1_2hTSB_2.fq.gz deltasbp_TSB_2h_1_R2.fastq.gz #1457dsbp2_2h_ ln -s ../F25A430000603_STAmsvaP/1457dsbp2_2hTSB/1457dsbp2_2hTSB_1.fq.gz deltasbp_TSB_2h_2_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457dsbp2_2hTSB/1457dsbp2_2hTSB_2.fq.gz deltasbp_TSB_2h_2_R2.fastq.gz #1457dsbp3_2h_ ln -s ../F25A430000603_STAmsvaP/1457dsbp3_2hTSB/1457dsbp3_2hTSB_1.fq.gz deltasbp_TSB_2h_3_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457dsbp3_2hTSB/1457dsbp3_2hTSB_2.fq.gz deltasbp_TSB_2h_3_R2.fastq.gz #1457dsbp1_4h_ ln -s ../F25A430000603_STAmsvaP/1457dsbp1_4hTSB/1457dsbp1_4hTSB_1.fq.gz deltasbp_TSB_4h_1_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457dsbp1_4hTSB/1457dsbp1_4hTSB_2.fq.gz deltasbp_TSB_4h_1_R2.fastq.gz #1457dsbp2_4h_ ln -s ../F25A430000603_STAmsvaP/1457dsbp2_4hTSB/1457dsbp2_4hTSB_1.fq.gz deltasbp_TSB_4h_2_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457dsbp2_4hTSB/1457dsbp2_4hTSB_2.fq.gz deltasbp_TSB_4h_2_R2.fastq.gz #1457dsbp3_4h_ ln -s ../F25A430000603_STAmsvaP/1457dsbp3_4hTSB/1457dsbp3_4hTSB_1.fq.gz deltasbp_TSB_4h_3_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457dsbp3_4hTSB/1457dsbp3_4hTSB_2.fq.gz deltasbp_TSB_4h_3_R2.fastq.gz #1457dsbp1_18h_ ln -s ../F25A430000603_STAmsvaP/1457dsbp118hTSB/1457dsbp118hTSB_1.fq.gz deltasbp_TSB_18h_1_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457dsbp118hTSB/1457dsbp118hTSB_2.fq.gz deltasbp_TSB_18h_1_R2.fastq.gz #1457dsbp2_18h_ ln -s ../F25A430000603_STAmsvaP/1457dsbp218hTSB/1457dsbp218hTSB_1.fq.gz deltasbp_TSB_18h_2_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457dsbp218hTSB/1457dsbp218hTSB_2.fq.gz deltasbp_TSB_18h_2_R2.fastq.gz #1457dsbp3_18h_ ln -s ../F25A430000603_STAmsvaP/1457dsbp318hTSB/1457dsbp318hTSB_1.fq.gz deltasbp_TSB_18h_3_R1.fastq.gz ln -s ../F25A430000603_STAmsvaP/1457dsbp318hTSB/1457dsbp318hTSB_2.fq.gz deltasbp_TSB_18h_3_R2.fastq.gz #END -
Preparing the directory trimmed
mkdir trimmed trimmed_unpaired; for sample_id in WT_MH_2h_1 WT_MH_2h_2 WT_MH_2h_3 WT_MH_4h_1 WT_MH_4h_2 WT_MH_4h_3 WT_MH_18h_1 WT_MH_18h_2 WT_MH_18h_3 WT_TSB_2h_1 WT_TSB_2h_2 WT_TSB_2h_3 WT_TSB_4h_1 WT_TSB_4h_2 WT_TSB_4h_3 WT_TSB_18h_1 WT_TSB_18h_2 WT_TSB_18h_3 deltasbp_MH_2h_1 deltasbp_MH_2h_2 deltasbp_MH_2h_3 deltasbp_MH_4h_1 deltasbp_MH_4h_2 deltasbp_MH_4h_3 deltasbp_MH_18h_1 deltasbp_MH_18h_2 deltasbp_TSB_2h_1 deltasbp_TSB_2h_2 deltasbp_TSB_2h_3 deltasbp_TSB_4h_1 deltasbp_TSB_4h_2 deltasbp_TSB_4h_3 deltasbp_TSB_18h_1 deltasbp_TSB_18h_2 deltasbp_TSB_18h_3; do java -jar /home/jhuang/Tools/Trimmomatic-0.36/trimmomatic-0.36.jar PE -threads 100 raw_data/${sample_id}_R1.fastq.gz raw_data/${sample_id}_R2.fastq.gz trimmed/${sample_id}_R1.fastq.gz trimmed_unpaired/${sample_id}_R1.fastq.gz trimmed/${sample_id}_R2.fastq.gz trimmed_unpaired/${sample_id}_R2.fastq.gz ILLUMINACLIP:/home/jhuang/Tools/Trimmomatic-0.36/adapters/TruSeq3-PE-2.fa:2:30:10:8:TRUE LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:36 AVGQUAL:20; done 2> trimmomatic_pe.log; done mv trimmed/*.fastq.gz . -
Preparing samplesheet.csv
sample,fastq_1,fastq_2,strandedness WT_MH_2h_1,WT_MH_2h_1_R1.fastq.gz,WT_MH_2h_1_R2.fastq.gz,auto ... -
nextflow run
#See an example: http://xgenes.com/article/article-content/157/prepare-virus-gtf-for-nextflow-run/ #docker pull nfcore/rnaseq ln -s /home/jhuang/Tools/nf-core-rnaseq-3.12.0/ rnaseq # -- DEBUG_1 (CDS --> exon in CP020463.gff) -- grep -P "\texon\t" CP020463.gff | sort | wc -l #=81 grep -P "cmsearch\texon\t" CP020463.gff | wc -l #=11 ignal recognition particle sRNA small typ, transfer-messenger RNA, 5S ribosomal RNA grep -P "Genbank\texon\t" CP020463.gff | wc -l #=12 16S and 23S ribosomal RNA grep -P "tRNAscan-SE\texon\t" CP020463.gff | wc -l #tRNA 58 grep -P "\tCDS\t" CP020463.gff | wc -l #3701-->2324 sed 's/\tCDS\t/\texon\t/g' CP020463.gff > CP020463_m.gff grep -P "\texon\t" CP020463_m.gff | sort | wc -l #3797-->2405 # -- NOTE that combination of 'CP020463_m.gff' and 'exon' in the command will result in ERROR, using 'transcript' instead in the command line! --gff "/home/jhuang/DATA/Data_Tam_RNAseq_2024/CP020463_m.gff" --featurecounts_feature_type 'transcript' # ---- SUCCESSFUL with directly downloaded gff3 and fasta from NCBI using docker after replacing 'CDS' with 'exon' ---- (host_env) /usr/local/bin/nextflow run rnaseq/main.nf --input samplesheet.csv --outdir results --fasta "/home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463.fasta" --gff "/home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_m.gff" -profile docker -resume --max_cpus 55 --max_memory 512.GB --max_time 2400.h --save_align_intermeds --save_unaligned --save_reference --aligner 'star_salmon' --gtf_group_features 'gene_id' --gtf_extra_attributes 'gene_name' --featurecounts_group_type 'gene_biotype' --featurecounts_feature_type 'transcript' # -- DEBUG_3: make sure the header of fasta is the same to the *_m.gff file, both are "CP020463.1" -
Import data and pca-plot
#mamba activate r_env #install.packages("ggfun") # Import the required libraries library("AnnotationDbi") library("clusterProfiler") library("ReactomePA") library(gplots) library(tximport) library(DESeq2) #library("org.Hs.eg.db") library(dplyr) library(tidyverse) #install.packages("devtools") #devtools::install_version("gtable", version = "0.3.0") library(gplots) library("RColorBrewer") #install.packages("ggrepel") library("ggrepel") # install.packages("openxlsx") library(openxlsx) library(EnhancedVolcano) library(DESeq2) library(edgeR) setwd("~/DATA/Data_Michelle_RNAseq_2025/results/star_salmon") # Define paths to your Salmon output quantification files files <- c( "deltasbp_MH_2h_r1" = "./deltasbp_MH_2h_1/quant.sf", "deltasbp_MH_2h_r2" = "./deltasbp_MH_2h_2/quant.sf", "deltasbp_MH_2h_r3" = "./deltasbp_MH_2h_3/quant.sf", "deltasbp_MH_4h_r1" = "./deltasbp_MH_4h_1/quant.sf", "deltasbp_MH_4h_r2" = "./deltasbp_MH_4h_2/quant.sf", "deltasbp_MH_4h_r3" = "./deltasbp_MH_4h_3/quant.sf", "deltasbp_MH_18h_r1" = "./deltasbp_MH_18h_1/quant.sf", "deltasbp_MH_18h_r2" = "./deltasbp_MH_18h_2/quant.sf", "deltasbp_TSB_2h_r1" = "./deltasbp_TSB_2h_1/quant.sf", "deltasbp_TSB_2h_r2" = "./deltasbp_TSB_2h_2/quant.sf", "deltasbp_TSB_2h_r3" = "./deltasbp_TSB_2h_3/quant.sf", "deltasbp_TSB_4h_r1" = "./deltasbp_TSB_4h_1/quant.sf", "deltasbp_TSB_4h_r2" = "./deltasbp_TSB_4h_2/quant.sf", "deltasbp_TSB_4h_r3" = "./deltasbp_TSB_4h_3/quant.sf", "deltasbp_TSB_18h_r1" = "./deltasbp_TSB_18h_1/quant.sf", "deltasbp_TSB_18h_r2" = "./deltasbp_TSB_18h_2/quant.sf", "deltasbp_TSB_18h_r3" = "./deltasbp_TSB_18h_3/quant.sf", "WT_MH_2h_r1" = "./WT_MH_2h_1/quant.sf", "WT_MH_2h_r2" = "./WT_MH_2h_2/quant.sf", "WT_MH_2h_r3" = "./WT_MH_2h_3/quant.sf", "WT_MH_4h_r1" = "./WT_MH_4h_1/quant.sf", "WT_MH_4h_r2" = "./WT_MH_4h_2/quant.sf", "WT_MH_4h_r3" = "./WT_MH_4h_3/quant.sf", "WT_MH_18h_r1" = "./WT_MH_18h_1/quant.sf", "WT_MH_18h_r2" = "./WT_MH_18h_2/quant.sf", "WT_MH_18h_r3" = "./WT_MH_18h_3/quant.sf", "WT_TSB_2h_r1" = "./WT_TSB_2h_1/quant.sf", "WT_TSB_2h_r2" = "./WT_TSB_2h_2/quant.sf", "WT_TSB_2h_r3" = "./WT_TSB_2h_3/quant.sf", "WT_TSB_4h_r1" = "./WT_TSB_4h_1/quant.sf", "WT_TSB_4h_r2" = "./WT_TSB_4h_2/quant.sf", "WT_TSB_4h_r3" = "./WT_TSB_4h_3/quant.sf", "WT_TSB_18h_r1" = "./WT_TSB_18h_1/quant.sf", "WT_TSB_18h_r2" = "./WT_TSB_18h_2/quant.sf", "WT_TSB_18h_r3" = "./WT_TSB_18h_3/quant.sf") # Import the transcript abundance data with tximport txi <- tximport(files, type = "salmon", txIn = TRUE, txOut = TRUE) # Define the replicates and condition of the samples replicate <- factor(c("r1","r2","r3", "r1","r2","r3", "r1","r2", "r1","r2","r3", "r1","r2","r3", "r1","r2","r3", "r1","r2","r3", "r1","r2","r3", "r1","r2","r3", "r1","r2","r3", "r1","r2","r3", "r1","r2","r3")) condition <- factor(c("deltasbp_MH_2h","deltasbp_MH_2h","deltasbp_MH_2h","deltasbp_MH_4h","deltasbp_MH_4h","deltasbp_MH_4h","deltasbp_MH_18h","deltasbp_MH_18h","deltasbp_TSB_2h","deltasbp_TSB_2h","deltasbp_TSB_2h","deltasbp_TSB_4h","deltasbp_TSB_4h","deltasbp_TSB_4h","deltasbp_TSB_18h","deltasbp_TSB_18h","deltasbp_TSB_18h","WT_MH_2h","WT_MH_2h","WT_MH_2h","WT_MH_4h","WT_MH_4h","WT_MH_4h","WT_MH_18h","WT_MH_18h","WT_MH_18h","WT_TSB_2h","WT_TSB_2h","WT_TSB_2h","WT_TSB_4h","WT_TSB_4h","WT_TSB_4h","WT_TSB_18h","WT_TSB_18h","WT_TSB_18h")) sample_table <- data.frame( condition = condition, replicate = replicate ) split_cond <- do.call(rbind, strsplit(as.character(condition), "_")) colnames(split_cond) <- c("strain", "media", "time") colData <- cbind(sample_table, split_cond) colData$strain <- factor(colData$strain) colData$media <- factor(colData$media) colData$time <- factor(colData$time) #colData$group <- factor(paste(colData$strain, colData$media, colData$time, sep = "_")) # Define the colData for DESeq2 #colData <- data.frame(condition=condition, row.names=names(files)) #grep "gene_name" ./results/genome/CP059040_m.gtf | wc -l #1701 #grep "gene_name" ./results/genome/CP020463_m.gtf | wc -l #50 # ------------------------ # 1️⃣ Setup and input files # ------------------------ # Read in transcript-to-gene mapping tx2gene <- read.table("salmon_tx2gene.tsv", header=FALSE, stringsAsFactors=FALSE) colnames(tx2gene) <- c("transcript_id", "gene_id", "gene_name") # Prepare tx2gene for gene-level summarization (remove gene_name if needed) tx2gene_geneonly <- tx2gene[, c("transcript_id", "gene_id")] # ------------------------------- # 2️⃣ Transcript-level counts # ------------------------------- # Create DESeqDataSet directly from tximport (transcript-level) dds_tx <- DESeqDataSetFromTximport(txi, colData=colData, design=~condition) write.csv(counts(dds_tx), file="transcript_counts.csv") # -------------------------------- # 3️⃣ Gene-level summarization # -------------------------------- # Re-import Salmon data summarized at gene level txi_gene <- tximport(files, type="salmon", tx2gene=tx2gene_geneonly, txOut=FALSE) # Create DESeqDataSet for gene-level counts #dds <- DESeqDataSetFromTximport(txi_gene, colData=colData, design=~condition+replicate) dds <- DESeqDataSetFromTximport(txi_gene, colData=colData, design=~condition) #dds <- DESeqDataSetFromTximport(txi, colData = colData, design = ~ time + media + strain + media:strain + strain:time) #或更简单地写为(推荐):dds <- DESeqDataSetFromTximport(txi, colData = colData, design = ~ time + media * strain) #dds <- DESeqDataSetFromTximport(txi, colData = colData, design = ~ strain * media * time) #~ strain * media * time 主效应 + 所有交互(推荐) ✅ #~ time + media * strain 主效应 + media:strain 交互 ⚠️ 有限制 # -------------------------------- # 4️⃣ Raw counts table (with gene names) # -------------------------------- # Extract raw gene-level counts counts_data <- as.data.frame(counts(dds, normalized=FALSE)) counts_data$gene_id <- rownames(counts_data) # Add gene names tx2gene_unique <- unique(tx2gene[, c("gene_id", "gene_name")]) counts_data <- merge(counts_data, tx2gene_unique, by="gene_id", all.x=TRUE) # Reorder columns: gene_id, gene_name, then counts count_cols <- setdiff(colnames(counts_data), c("gene_id", "gene_name")) counts_data <- counts_data[, c("gene_id", "gene_name", count_cols)] # -------------------------------- # 5️⃣ Calculate CPM # -------------------------------- library(edgeR) library(openxlsx) # Prepare count matrix for CPM calculation count_matrix <- as.matrix(counts_data[, !(colnames(counts_data) %in% c("gene_id", "gene_name"))]) # Calculate CPM #cpm_matrix <- cpm(count_matrix, normalized.lib.sizes=FALSE) total_counts <- colSums(count_matrix) cpm_matrix <- t(t(count_matrix) / total_counts) * 1e6 cpm_matrix <- as.data.frame(cpm_matrix) # Add gene_id and gene_name back to CPM table cpm_counts <- cbind(counts_data[, c("gene_id", "gene_name")], cpm_matrix) # -------------------------------- # 6️⃣ Save outputs # -------------------------------- write.csv(counts_data, "gene_raw_counts.csv", row.names=FALSE) write.xlsx(counts_data, "gene_raw_counts.xlsx", row.names=FALSE) write.xlsx(cpm_counts, "gene_cpm_counts.xlsx", row.names=FALSE) # -- (Optional) Save the rlog-transformed counts -- dim(counts(dds)) head(counts(dds), 10) rld <- rlogTransformation(dds) rlog_counts <- assay(rld) write.xlsx(as.data.frame(rlog_counts), "gene_rlog_transformed_counts.xlsx") # -- pca -- png("pca2.png", 1200, 800) plotPCA(rld, intgroup=c("condition")) dev.off() # -- heatmap -- png("heatmap2.png", 1200, 800) distsRL <- dist(t(assay(rld))) mat <- as.matrix(distsRL) hc <- hclust(distsRL) hmcol <- colorRampPalette(brewer.pal(9,"GnBu"))(100) heatmap.2(mat, Rowv=as.dendrogram(hc),symm=TRUE, trace="none",col = rev(hmcol), margin=c(13, 13)) dev.off() # -- pca_media_strain -- png("pca_media.png", 1200, 800) plotPCA(rld, intgroup=c("media")) dev.off() png("pca_strain.png", 1200, 800) plotPCA(rld, intgroup=c("strain")) dev.off() png("pca_time.png", 1200, 800) plotPCA(rld, intgroup=c("time")) dev.off() -
(Optional; ERROR-->need to be debugged!) ) estimate size factors and dispersion values.
#Size Factors: These are used to normalize the read counts across different samples. The size factor for a sample accounts for differences in sequencing depth (i.e., the total number of reads) and other technical biases between samples. After normalization with size factors, the counts should be comparable across samples. Size factors are usually calculated in a way that they reflect the median or mean ratio of gene expression levels between samples, assuming that most genes are not differentially expressed. #Dispersion: This refers to the variability or spread of gene expression measurements. In RNA-seq data analysis, each gene has its own dispersion value, which reflects how much the counts for that gene vary between different samples, more than what would be expected just due to the Poisson variation inherent in counting. Dispersion is important for accurately modeling the data and for detecting differentially expressed genes. #So in summary, size factors are specific to samples (used to make counts comparable across samples), and dispersion values are specific to genes (reflecting variability in gene expression). sizeFactors(dds) #NULL # Estimate size factors dds <- estimateSizeFactors(dds) # Estimate dispersions dds <- estimateDispersions(dds) #> sizeFactors(dds) #control_r1 control_r2 HSV.d2_r1 HSV.d2_r2 HSV.d4_r1 HSV.d4_r2 HSV.d6_r1 #2.3282468 2.0251928 1.8036883 1.3767551 0.9341929 1.0911693 0.5454526 #HSV.d6_r2 HSV.d8_r1 HSV.d8_r2 #0.4604461 0.5799834 0.6803681 # (DEBUG) If avgTxLength is Necessary #To simplify the computation and ensure sizeFactors are calculated: assays(dds)$avgTxLength <- NULL dds <- estimateSizeFactors(dds) sizeFactors(dds) #If you want to retain avgTxLength but suspect it is causing issues, you can explicitly instruct DESeq2 to compute size factors without correcting for library size with average transcript lengths: dds <- estimateSizeFactors(dds, controlGenes = NULL, use = FALSE) sizeFactors(dds) # If alone with virus data, the following BUG occured: #Still NULL --> BUG --> using manual calculation method for sizeFactor calculation! HeLa_TO_r1 HeLa_TO_r2 0.9978755 1.1092227 data.frame(genes = rownames(dds), dispersions = dispersions(dds)) #Given the raw counts, the control_r1 and control_r2 samples seem to have a much lower sequencing depth (total read count) than the other samples. Therefore, when normalization methods are applied, the normalization factors for these control samples will be relatively high, boosting the normalized counts. 1/0.9978755=1.002129023 1/1.1092227= #bamCoverage --bam ../markDuplicates/${sample}Aligned.sortedByCoord.out.bam -o ${sample}_norm.bw --binSize 10 --scaleFactor --effectiveGenomeSize 2864785220 bamCoverage --bam ../markDuplicates/HeLa_TO_r1Aligned.sortedByCoord.out.markDups.bam -o HeLa_TO_r1.bw --binSize 10 --scaleFactor 1.002129023 --effectiveGenomeSize 2864785220 bamCoverage --bam ../markDuplicates/HeLa_TO_r2Aligned.sortedByCoord.out.markDups.bam -o HeLa_TO_r2.bw --binSize 10 --scaleFactor 0.901532217 --effectiveGenomeSize 2864785220 raw_counts <- counts(dds) normalized_counts <- counts(dds, normalized=TRUE) #write.table(raw_counts, file="raw_counts.txt", sep="\t", quote=F, col.names=NA) #write.table(normalized_counts, file="normalized_counts.txt", sep="\t", quote=F, col.names=NA) #convert bam to bigwig using deepTools by feeding inverse of DESeq’s size Factor estimSf <- function (cds){ # Get the count matrix cts <- counts(cds) # Compute the geometric mean geomMean <- function(x) prod(x)^(1/length(x)) # Compute the geometric mean over the line gm.mean <- apply(cts, 1, geomMean) # Zero values are set to NA (avoid subsequentcdsdivision by 0) gm.mean[gm.mean == 0] <- NA # Divide each line by its corresponding geometric mean # sweep(x, MARGIN, STATS, FUN = "-", check.margin = TRUE, ...) # MARGIN: 1 or 2 (line or columns) # STATS: a vector of length nrow(x) or ncol(x), depending on MARGIN # FUN: the function to be applied cts <- sweep(cts, 1, gm.mean, FUN="/") # Compute the median over the columns med <- apply(cts, 2, median, na.rm=TRUE) # Return the scaling factor return(med) } #https://dputhier.github.io/ASG/practicals/rnaseq_diff_Snf2/rnaseq_diff_Snf2.html #http://bioconductor.org/packages/devel/bioc/vignettes/DESeq2/inst/doc/DESeq2.html#data-transformations-and-visualization #https://hbctraining.github.io/DGE_workshop/lessons/02_DGE_count_normalization.html #https://hbctraining.github.io/DGE_workshop/lessons/04_DGE_DESeq2_analysis.html #https://genviz.org/module-04-expression/0004/02/01/DifferentialExpression/ #DESeq2’s median of ratios [1] #EdgeR’s trimmed mean of M values (TMM) [2] #http://www.nathalievialaneix.eu/doc/html/TP1_normalization.html #very good website! test_normcount <- sweep(raw_counts, 2, sizeFactors(dds), "/") sum(test_normcount != normalized_counts) -
Select the differentially expressed genes
#https://galaxyproject.eu/posts/2020/08/22/three-steps-to-galaxify-your-tool/ #https://www.biostars.org/p/282295/ #https://www.biostars.org/p/335751/ dds$condition [1] deltasbp_MH_2h deltasbp_MH_2h deltasbp_MH_2h deltasbp_MH_4h [5] deltasbp_MH_4h deltasbp_MH_4h deltasbp_MH_18h deltasbp_MH_18h [9] deltasbp_TSB_2h deltasbp_TSB_2h deltasbp_TSB_2h deltasbp_TSB_4h [13] deltasbp_TSB_4h deltasbp_TSB_4h deltasbp_TSB_18h deltasbp_TSB_18h [17] deltasbp_TSB_18h WT_MH_2h WT_MH_2h WT_MH_2h [21] WT_MH_4h WT_MH_4h WT_MH_4h WT_MH_18h [25] WT_MH_18h WT_MH_18h WT_TSB_2h WT_TSB_2h [29] WT_TSB_2h WT_TSB_4h WT_TSB_4h WT_TSB_4h [33] WT_TSB_18h WT_TSB_18h WT_TSB_18h 12 Levels: deltasbp_MH_18h deltasbp_MH_2h deltasbp_MH_4h ... WT_TSB_4h #CONSOLE: mkdir star_salmon/degenes setwd("degenes") # 确保因子顺序(可选) colData$strain <- relevel(factor(colData$strain), ref = "WT") colData$media <- relevel(factor(colData$media), ref = "TSB") colData$time <- relevel(factor(colData$time), ref = "2h") dds <- DESeqDataSetFromTximport(txi, colData, design = ~ strain * media * time) dds <- DESeq(dds, betaPrior = FALSE) resultsNames(dds) #[1] "Intercept" "strain_deltasbp_vs_WT" #[3] "media_MH_vs_TSB" "time_18h_vs_2h" #[5] "time_4h_vs_2h" "straindeltasbp.mediaMH" #[7] "straindeltasbp.time18h" "straindeltasbp.time4h" #[9] "mediaMH.time18h" "mediaMH.time4h" #[11] "straindeltasbp.mediaMH.time18h" "straindeltasbp.mediaMH.time4h" 🔹 Main effects for each factor: 表达量 ▲ │ ┌────── WT-TSB │ / │ / ┌────── WT-MH │ / / │ / / ┌────── deltasbp-TSB │ / / / │ / / / ┌────── deltasbp-MH └──────────────────────────────▶ 时间(2h, 4h, 18h) * strain_deltasbp_vs_WT * media_MH_vs_TSB * time_18h_vs_2h * time_4h_vs_2h 🔹 两因素交互作用(Two-way interactions) 这些项表示两个实验因素(如菌株、培养基、时间)之间的组合效应——也就是说,其中一个因素的影响取决于另一个因素的水平。 表达量 ▲ │ │ WT ────────┐ │ └─↘ │ ↘ │ deltasbp ←←←← 显著交互(方向/幅度不同) └──────────────────────────────▶ 时间 straindeltasbp.mediaMH 表示 菌株(strain)和培养基(media)之间的交互作用。 ➤ 这意味着:deltasbp 这个突变菌株在 MH 培养基中的表现与它在 TSB 中的不同,不能仅通过菌株和培养基的单独效应来解释。 straindeltasbp.time18h 表示 菌株(strain)和时间(time, 18h)之间的交互作用。 ➤ 即:突变菌株在 18 小时时的表达变化不只是菌株效应或时间效应的简单相加,而有协同作用。 straindeltasbp.time4h 同上,是菌株和时间(4h)之间的交互作用。 mediaMH.time18h 表示 培养基(MH)与时间(18h)之间的交互作用。 ➤ 即:在 MH 培养基中,18 小时时的表达水平与在其他时间点(例如 2h)不同,且该变化不完全可以用时间和培养基各自单独的效应来解释。 mediaMH.time4h 与上面类似,是 MH 培养基与 4 小时之间的交互作用。 🔹 三因素交互作用(Three-way interactions) 三因素交互作用表示:菌株、培养基和时间这三个因素在一起时,会产生一个新的效应,这种效应无法通过任何两个因素的组合来完全解释。 表达量(TSB) ▲ │ │ WT ──────→→ │ deltasbp ─────→→ └────────────────────────▶ 时间(2h, 4h, 18h) 表达量(MH) ▲ │ │ WT ──────→→ │ deltasbp ─────⬈⬈⬈⬈⬈⬈⬈ └────────────────────────▶ 时间(2h, 4h, 18h) straindeltasbp.mediaMH.time18h 表示 菌株 × 培养基 × 时间(18h) 三者之间的交互作用。 ➤ 即:突变菌株在 MH 培养基下的 18 小时表达模式,与其他组合(比如 WT 在 MH 培养基下,或者在 TSB 下)都不相同。 straindeltasbp.mediaMH.time4h 同上,只是观察的是 4 小时下的三因素交互效应。 ✅ 总结: 交互作用项的存在意味着你不能仅通过单个变量(如菌株、时间或培养基)的影响来解释基因表达的变化,必须同时考虑它们之间的组合关系。在 DESeq2 模型中,这些交互项的显著性可以揭示特定条件下是否有特异的调控行为。 # 提取 strain 的主效应: up 2, down 16 contrast <- "strain_deltasbp_vs_WT" res = results(dds, name=contrast) res <- res[!is.na(res$log2FoldChange),] res_df <- as.data.frame(res) write.csv(as.data.frame(res_df[order(res_df$pvalue),]), file = paste(contrast, "all.txt", sep="-")) up <- subset(res_df, padj<=0.05 & log2FoldChange>=2) down <- subset(res_df, padj<=0.05 & log2FoldChange<=-2) write.csv(as.data.frame(up[order(up$log2FoldChange,decreasing=TRUE),]), file = paste(contrast, "up.txt", sep="-")) write.csv(as.data.frame(down[order(abs(down$log2FoldChange),decreasing=TRUE),]), file = paste(contrast, "down.txt", sep="-")) # 提取 media 的主效应: up 76; down 128 contrast <- "media_MH_vs_TSB" res = results(dds, name=contrast) res <- res[!is.na(res$log2FoldChange),] res_df <- as.data.frame(res) write.csv(as.data.frame(res_df[order(res_df$pvalue),]), file = paste(contrast, "all.txt", sep="-")) up <- subset(res_df, padj<=0.05 & log2FoldChange>=2) down <- subset(res_df, padj<=0.05 & log2FoldChange<=-2) write.csv(as.data.frame(up[order(up$log2FoldChange,decreasing=TRUE),]), file = paste(contrast, "up.txt", sep="-")) write.csv(as.data.frame(down[order(abs(down$log2FoldChange),decreasing=TRUE),]), file = paste(contrast, "down.txt", sep="-")) # 提取 time 的主效应 up 228, down 98; up 17, down 2 contrast <- "time_18h_vs_2h" res = results(dds, name=contrast) res <- res[!is.na(res$log2FoldChange),] res_df <- as.data.frame(res) write.csv(as.data.frame(res_df[order(res_df$pvalue),]), file = paste(contrast, "all.txt", sep="-")) up <- subset(res_df, padj<=0.05 & log2FoldChange>=2) down <- subset(res_df, padj<=0.05 & log2FoldChange<=-2) write.csv(as.data.frame(up[order(up$log2FoldChange,decreasing=TRUE),]), file = paste(contrast, "up.txt", sep="-")) write.csv(as.data.frame(down[order(abs(down$log2FoldChange),decreasing=TRUE),]), file = paste(contrast, "down.txt", sep="-")) contrast <- "time_4h_vs_2h" res = results(dds, name=contrast) res <- res[!is.na(res$log2FoldChange),] res_df <- as.data.frame(res) write.csv(as.data.frame(res_df[order(res_df$pvalue),]), file = paste(contrast, "all.txt", sep="-")) up <- subset(res_df, padj<=0.05 & log2FoldChange>=2) down <- subset(res_df, padj<=0.05 & log2FoldChange<=-2) write.csv(as.data.frame(up[order(up$log2FoldChange,decreasing=TRUE),]), file = paste(contrast, "up.txt", sep="-")) write.csv(as.data.frame(down[order(abs(down$log2FoldChange),decreasing=TRUE),]), file = paste(contrast, "down.txt", sep="-")) #1.) delta sbp 2h TSB vs WT 2h TSB #2.) delta sbp 4h TSB vs WT 4h TSB #3.) delta sbp 18h TSB vs WT 18h TSB #4.) delta sbp 2h MH vs WT 2h MH #5.) delta sbp 4h MH vs WT 4h MH #6.) delta sbp 18h MH vs WT 18h MH #---- relevel to control ---- #design=~condition+replicate dds <- DESeqDataSetFromTximport(txi, colData, design = ~ condition) dds$condition <- relevel(dds$condition, "WT_TSB_2h") dds = DESeq(dds, betaPrior=FALSE) resultsNames(dds) clist <- c("deltasbp_TSB_2h_vs_WT_TSB_2h") dds$condition <- relevel(dds$condition, "WT_TSB_4h") dds = DESeq(dds, betaPrior=FALSE) resultsNames(dds) clist <- c("deltasbp_TSB_4h_vs_WT_TSB_4h") dds$condition <- relevel(dds$condition, "WT_TSB_18h") dds = DESeq(dds, betaPrior=FALSE) resultsNames(dds) clist <- c("deltasbp_TSB_18h_vs_WT_TSB_18h") dds$condition <- relevel(dds$condition, "WT_MH_2h") dds = DESeq(dds, betaPrior=FALSE) resultsNames(dds) clist <- c("deltasbp_MH_2h_vs_WT_MH_2h") dds$condition <- relevel(dds$condition, "WT_MH_4h") dds = DESeq(dds, betaPrior=FALSE) resultsNames(dds) clist <- c("deltasbp_MH_4h_vs_WT_MH_4h") dds$condition <- relevel(dds$condition, "WT_MH_18h") dds = DESeq(dds, betaPrior=FALSE) resultsNames(dds) clist <- c("deltasbp_MH_18h_vs_WT_MH_18h") for (i in clist) { contrast = paste("condition", i, sep="_") #for_Mac_vs_LB contrast = paste("media", i, sep="_") res = results(dds, name=contrast) res <- res[!is.na(res$log2FoldChange),] res_df <- as.data.frame(res) write.csv(as.data.frame(res_df[order(res_df$pvalue),]), file = paste(i, "all.txt", sep="-")) #res$log2FoldChange < -2 & res$padj < 1e-2 up <- subset(res_df, padj<=0.01 & log2FoldChange>=2) down <- subset(res_df, padj<=0.01 & log2FoldChange<=-2) write.csv(as.data.frame(up[order(up$log2FoldChange,decreasing=TRUE),]), file = paste(i, "up.txt", sep="-")) write.csv(as.data.frame(down[order(abs(down$log2FoldChange),decreasing=TRUE),]), file = paste(i, "down.txt", sep="-")) } # -- Under host-env (mamba activate plot-numpy1) -- grep -P "\tgene\t" CP020463.gff > CP020463_gene.gff python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_TSB_2h_vs_WT_TSB_2h-all.txt deltasbp_TSB_2h_vs_WT_TSB_2h-all.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_TSB_2h_vs_WT_TSB_2h-up.txt deltasbp_TSB_2h_vs_WT_TSB_2h-up.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_TSB_2h_vs_WT_TSB_2h-down.txt deltasbp_TSB_2h_vs_WT_TSB_2h-down.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_TSB_4h_vs_WT_TSB_4h-all.txt deltasbp_TSB_4h_vs_WT_TSB_4h-all.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_TSB_4h_vs_WT_TSB_4h-up.txt deltasbp_TSB_4h_vs_WT_TSB_4h-up.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_TSB_4h_vs_WT_TSB_4h-down.txt deltasbp_TSB_4h_vs_WT_TSB_4h-down.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_TSB_18h_vs_WT_TSB_18h-all.txt deltasbp_TSB_18h_vs_WT_TSB_18h-all.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_TSB_18h_vs_WT_TSB_18h-up.txt deltasbp_TSB_18h_vs_WT_TSB_18h-up.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_TSB_18h_vs_WT_TSB_18h-down.txt deltasbp_TSB_18h_vs_WT_TSB_18h-down.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_MH_2h_vs_WT_MH_2h-all.txt deltasbp_MH_2h_vs_WT_MH_2h-all.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_MH_2h_vs_WT_MH_2h-up.txt deltasbp_MH_2h_vs_WT_MH_2h-up.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_MH_2h_vs_WT_MH_2h-down.txt deltasbp_MH_2h_vs_WT_MH_2h-down.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_MH_4h_vs_WT_MH_4h-all.txt deltasbp_MH_4h_vs_WT_MH_4h-all.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_MH_4h_vs_WT_MH_4h-up.txt deltasbp_MH_4h_vs_WT_MH_4h-up.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_MH_4h_vs_WT_MH_4h-down.txt deltasbp_MH_4h_vs_WT_MH_4h-down.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_MH_18h_vs_WT_MH_18h-all.txt deltasbp_MH_18h_vs_WT_MH_18h-all.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_MH_18h_vs_WT_MH_18h-up.txt deltasbp_MH_18h_vs_WT_MH_18h-up.csv python3 ~/Scripts/replace_gene_names.py /home/jhuang/DATA/Data_Michelle_RNAseq_2025/CP020463_gene.gff deltasbp_MH_18h_vs_WT_MH_18h-down.txt deltasbp_MH_18h_vs_WT_MH_18h-down.csv # ---- delta sbp TSB 2h vs WT TSB 2h ---- res <- read.csv("deltasbp_TSB_2h_vs_WT_TSB_2h-all.csv") # Replace empty GeneName with modified GeneID res$GeneName <- ifelse( res$GeneName == "" | is.na(res$GeneName), gsub("gene-", "", res$GeneID), res$GeneName ) duplicated_genes <- res[duplicated(res$GeneName), "GeneName"] #print(duplicated_genes) # [1] "bfr" "lipA" "ahpF" "pcaF" "alr" "pcaD" "cydB" "lpdA" "pgaC" "ppk1" #[11] "pcaF" "tuf" "galE" "murI" "yccS" "rrf" "rrf" "arsB" "ptsP" "umuD" #[21] "map" "pgaB" "rrf" "rrf" "rrf" "pgaD" "uraH" "benE" #res[res$GeneName == "bfr", ] #1st_strategy First occurrence is kept and Subsequent duplicates are removed #res <- res[!duplicated(res$GeneName), ] #2nd_strategy keep the row with the smallest padj value for each GeneName res <- res %>% group_by(GeneName) %>% slice_min(padj, with_ties = FALSE) %>% ungroup() res <- as.data.frame(res) # Sort res first by padj (ascending) and then by log2FoldChange (descending) res <- res[order(res$padj, -res$log2FoldChange), ] # Assuming res is your dataframe and already processed # Filter up-regulated genes: log2FoldChange > 2 and padj < 1e-2 up_regulated <- res[res$log2FoldChange > 2 & res$padj < 1e-2, ] # Filter down-regulated genes: log2FoldChange < -2 and padj < 1e-2 down_regulated <- res[res$log2FoldChange < -2 & res$padj < 1e-2, ] # Create a new workbook wb <- createWorkbook() # Add the complete dataset as the first sheet addWorksheet(wb, "Complete_Data") writeData(wb, "Complete_Data", res) # Add the up-regulated genes as the second sheet addWorksheet(wb, "Up_Regulated") writeData(wb, "Up_Regulated", up_regulated) # Add the down-regulated genes as the third sheet addWorksheet(wb, "Down_Regulated") writeData(wb, "Down_Regulated", down_regulated) # Save the workbook to a file saveWorkbook(wb, "Gene_Expression_Δsbp_TSB_2h_vs_WT_TSB_2h.xlsx", overwrite = TRUE) # Set the 'GeneName' column as row.names rownames(res) <- res$GeneName # Drop the 'GeneName' column since it's now the row names res$GeneName <- NULL head(res) ## Ensure the data frame matches the expected format ## For example, it should have columns: log2FoldChange, padj, etc. #res <- as.data.frame(res) ## Remove rows with NA in log2FoldChange (if needed) #res <- res[!is.na(res$log2FoldChange),] # Replace padj = 0 with a small value #NO_SUCH_RECORDS: res$padj[res$padj == 0] <- 1e-150 #library(EnhancedVolcano) # Assuming res is already sorted and processed png("Δsbp_TSB_2h_vs_WT_TSB_2h.png", width=1200, height=1200) #max.overlaps = 10 EnhancedVolcano(res, lab = rownames(res), x = 'log2FoldChange', y = 'padj', pCutoff = 1e-2, FCcutoff = 2, title = '', subtitleLabSize = 18, pointSize = 3.0, labSize = 5.0, colAlpha = 1, legendIconSize = 4.0, drawConnectors = TRUE, widthConnectors = 0.5, colConnectors = 'black', subtitle = expression("Δsbp TSB 2h versus WT TSB 2h")) dev.off() # ---- delta sbp TSB 4h vs WT TSB 4h ---- res <- read.csv("deltasbp_TSB_4h_vs_WT_TSB_4h-all.csv") # Replace empty GeneName with modified GeneID res$GeneName <- ifelse( res$GeneName == "" | is.na(res$GeneName), gsub("gene-", "", res$GeneID), res$GeneName ) duplicated_genes <- res[duplicated(res$GeneName), "GeneName"] res <- res %>% group_by(GeneName) %>% slice_min(padj, with_ties = FALSE) %>% ungroup() res <- as.data.frame(res) # Sort res first by padj (ascending) and then by log2FoldChange (descending) res <- res[order(res$padj, -res$log2FoldChange), ] # Assuming res is your dataframe and already processed # Filter up-regulated genes: log2FoldChange > 2 and padj < 1e-2 up_regulated <- res[res$log2FoldChange > 2 & res$padj < 1e-2, ] # Filter down-regulated genes: log2FoldChange < -2 and padj < 1e-2 down_regulated <- res[res$log2FoldChange < -2 & res$padj < 1e-2, ] # Create a new workbook wb <- createWorkbook() # Add the complete dataset as the first sheet addWorksheet(wb, "Complete_Data") writeData(wb, "Complete_Data", res) # Add the up-regulated genes as the second sheet addWorksheet(wb, "Up_Regulated") writeData(wb, "Up_Regulated", up_regulated) # Add the down-regulated genes as the third sheet addWorksheet(wb, "Down_Regulated") writeData(wb, "Down_Regulated", down_regulated) # Save the workbook to a file saveWorkbook(wb, "Gene_Expression_Δsbp_TSB_4h_vs_WT_TSB_4h.xlsx", overwrite = TRUE) # Set the 'GeneName' column as row.names rownames(res) <- res$GeneName # Drop the 'GeneName' column since it's now the row names res$GeneName <- NULL head(res) #library(EnhancedVolcano) # Assuming res is already sorted and processed png("Δsbp_TSB_4h_vs_WT_TSB_4h.png", width=1200, height=1200) #max.overlaps = 10 EnhancedVolcano(res, lab = rownames(res), x = 'log2FoldChange', y = 'padj', pCutoff = 1e-2, FCcutoff = 2, title = '', subtitleLabSize = 18, pointSize = 3.0, labSize = 5.0, colAlpha = 1, legendIconSize = 4.0, drawConnectors = TRUE, widthConnectors = 0.5, colConnectors = 'black', subtitle = expression("Δsbp TSB 4h versus WT TSB 4h")) dev.off() # ---- delta sbp TSB 18h vs WT TSB 18h ---- res <- read.csv("deltasbp_TSB_18h_vs_WT_TSB_18h-all.csv") # Replace empty GeneName with modified GeneID res$GeneName <- ifelse( res$GeneName == "" | is.na(res$GeneName), gsub("gene-", "", res$GeneID), res$GeneName ) duplicated_genes <- res[duplicated(res$GeneName), "GeneName"] res <- res %>% group_by(GeneName) %>% slice_min(padj, with_ties = FALSE) %>% ungroup() res <- as.data.frame(res) # Sort res first by padj (ascending) and then by log2FoldChange (descending) res <- res[order(res$padj, -res$log2FoldChange), ] # Assuming res is your dataframe and already processed # Filter up-regulated genes: log2FoldChange > 2 and padj < 1e-2 up_regulated <- res[res$log2FoldChange > 2 & res$padj < 1e-2, ] # Filter down-regulated genes: log2FoldChange < -2 and padj < 1e-2 down_regulated <- res[res$log2FoldChange < -2 & res$padj < 1e-2, ] # Create a new workbook wb <- createWorkbook() # Add the complete dataset as the first sheet addWorksheet(wb, "Complete_Data") writeData(wb, "Complete_Data", res) # Add the up-regulated genes as the second sheet addWorksheet(wb, "Up_Regulated") writeData(wb, "Up_Regulated", up_regulated) # Add the down-regulated genes as the third sheet addWorksheet(wb, "Down_Regulated") writeData(wb, "Down_Regulated", down_regulated) # Save the workbook to a file saveWorkbook(wb, "Gene_Expression_Δsbp_TSB_18h_vs_WT_TSB_18h.xlsx", overwrite = TRUE) # Set the 'GeneName' column as row.names rownames(res) <- res$GeneName # Drop the 'GeneName' column since it's now the row names res$GeneName <- NULL head(res) #library(EnhancedVolcano) # Assuming res is already sorted and processed png("Δsbp_TSB_18h_vs_WT_TSB_18h.png", width=1200, height=1200) #max.overlaps = 10 EnhancedVolcano(res, lab = rownames(res), x = 'log2FoldChange', y = 'padj', pCutoff = 1e-2, FCcutoff = 2, title = '', subtitleLabSize = 18, pointSize = 3.0, labSize = 5.0, colAlpha = 1, legendIconSize = 4.0, drawConnectors = TRUE, widthConnectors = 0.5, colConnectors = 'black', subtitle = expression("Δsbp TSB 18h versus WT TSB 18h")) dev.off() # ---- delta sbp MH 2h vs WT MH 2h ---- res <- read.csv("deltasbp_MH_2h_vs_WT_MH_2h-all.csv") # Replace empty GeneName with modified GeneID res$GeneName <- ifelse( res$GeneName == "" | is.na(res$GeneName), gsub("gene-", "", res$GeneID), res$GeneName ) duplicated_genes <- res[duplicated(res$GeneName), "GeneName"] #print(duplicated_genes) # [1] "bfr" "lipA" "ahpF" "pcaF" "alr" "pcaD" "cydB" "lpdA" "pgaC" "ppk1" #[11] "pcaF" "tuf" "galE" "murI" "yccS" "rrf" "rrf" "arsB" "ptsP" "umuD" #[21] "map" "pgaB" "rrf" "rrf" "rrf" "pgaD" "uraH" "benE" #res[res$GeneName == "bfr", ] #1st_strategy First occurrence is kept and Subsequent duplicates are removed #res <- res[!duplicated(res$GeneName), ] #2nd_strategy keep the row with the smallest padj value for each GeneName res <- res %>% group_by(GeneName) %>% slice_min(padj, with_ties = FALSE) %>% ungroup() res <- as.data.frame(res) # Sort res first by padj (ascending) and then by log2FoldChange (descending) res <- res[order(res$padj, -res$log2FoldChange), ] # Assuming res is your dataframe and already processed # Filter up-regulated genes: log2FoldChange > 2 and padj < 1e-2 up_regulated <- res[res$log2FoldChange > 2 & res$padj < 1e-2, ] # Filter down-regulated genes: log2FoldChange < -2 and padj < 1e-2 down_regulated <- res[res$log2FoldChange < -2 & res$padj < 1e-2, ] # Create a new workbook wb <- createWorkbook() # Add the complete dataset as the first sheet addWorksheet(wb, "Complete_Data") writeData(wb, "Complete_Data", res) # Add the up-regulated genes as the second sheet addWorksheet(wb, "Up_Regulated") writeData(wb, "Up_Regulated", up_regulated) # Add the down-regulated genes as the third sheet addWorksheet(wb, "Down_Regulated") writeData(wb, "Down_Regulated", down_regulated) # Save the workbook to a file saveWorkbook(wb, "Gene_Expression_Δsbp_MH_2h_vs_WT_MH_2h.xlsx", overwrite = TRUE) # Set the 'GeneName' column as row.names rownames(res) <- res$GeneName # Drop the 'GeneName' column since it's now the row names res$GeneName <- NULL head(res) ## Ensure the data frame matches the expected format ## For example, it should have columns: log2FoldChange, padj, etc. #res <- as.data.frame(res) ## Remove rows with NA in log2FoldChange (if needed) #res <- res[!is.na(res$log2FoldChange),] # Replace padj = 0 with a small value #NO_SUCH_RECORDS: res$padj[res$padj == 0] <- 1e-150 #library(EnhancedVolcano) # Assuming res is already sorted and processed png("Δsbp_MH_2h_vs_WT_MH_2h.png", width=1200, height=1200) #max.overlaps = 10 EnhancedVolcano(res, lab = rownames(res), x = 'log2FoldChange', y = 'padj', pCutoff = 1e-2, FCcutoff = 2, title = '', subtitleLabSize = 18, pointSize = 3.0, labSize = 5.0, colAlpha = 1, legendIconSize = 4.0, drawConnectors = TRUE, widthConnectors = 0.5, colConnectors = 'black', subtitle = expression("Δsbp MH 2h versus WT MH 2h")) dev.off() # ---- delta sbp MH 4h vs WT MH 4h ---- res <- read.csv("deltasbp_MH_4h_vs_WT_MH_4h-all.csv") # Replace empty GeneName with modified GeneID res$GeneName <- ifelse( res$GeneName == "" | is.na(res$GeneName), gsub("gene-", "", res$GeneID), res$GeneName ) duplicated_genes <- res[duplicated(res$GeneName), "GeneName"] res <- res %>% group_by(GeneName) %>% slice_min(padj, with_ties = FALSE) %>% ungroup() res <- as.data.frame(res) # Sort res first by padj (ascending) and then by log2FoldChange (descending) res <- res[order(res$padj, -res$log2FoldChange), ] # Assuming res is your dataframe and already processed # Filter up-regulated genes: log2FoldChange > 2 and padj < 1e-2 up_regulated <- res[res$log2FoldChange > 2 & res$padj < 1e-2, ] # Filter down-regulated genes: log2FoldChange < -2 and padj < 1e-2 down_regulated <- res[res$log2FoldChange < -2 & res$padj < 1e-2, ] # Create a new workbook wb <- createWorkbook() # Add the complete dataset as the first sheet addWorksheet(wb, "Complete_Data") writeData(wb, "Complete_Data", res) # Add the up-regulated genes as the second sheet addWorksheet(wb, "Up_Regulated") writeData(wb, "Up_Regulated", up_regulated) # Add the down-regulated genes as the third sheet addWorksheet(wb, "Down_Regulated") writeData(wb, "Down_Regulated", down_regulated) # Save the workbook to a file saveWorkbook(wb, "Gene_Expression_Δsbp_MH_4h_vs_WT_MH_4h.xlsx", overwrite = TRUE) # Set the 'GeneName' column as row.names rownames(res) <- res$GeneName # Drop the 'GeneName' column since it's now the row names res$GeneName <- NULL head(res) #library(EnhancedVolcano) # Assuming res is already sorted and processed png("Δsbp_MH_4h_vs_WT_MH_4h.png", width=1200, height=1200) #max.overlaps = 10 EnhancedVolcano(res, lab = rownames(res), x = 'log2FoldChange', y = 'padj', pCutoff = 1e-2, FCcutoff = 2, title = '', subtitleLabSize = 18, pointSize = 3.0, labSize = 5.0, colAlpha = 1, legendIconSize = 4.0, drawConnectors = TRUE, widthConnectors = 0.5, colConnectors = 'black', subtitle = expression("Δsbp MH 4h versus WT MH 4h")) dev.off() # ---- delta sbp MH 18h vs WT MH 18h ---- res <- read.csv("deltasbp_MH_18h_vs_WT_MH_18h-all.csv") # Replace empty GeneName with modified GeneID res$GeneName <- ifelse( res$GeneName == "" | is.na(res$GeneName), gsub("gene-", "", res$GeneID), res$GeneName ) duplicated_genes <- res[duplicated(res$GeneName), "GeneName"] res <- res %>% group_by(GeneName) %>% slice_min(padj, with_ties = FALSE) %>% ungroup() res <- as.data.frame(res) # Sort res first by padj (ascending) and then by log2FoldChange (descending) res <- res[order(res$padj, -res$log2FoldChange), ] # Assuming res is your dataframe and already processed # Filter up-regulated genes: log2FoldChange > 2 and padj < 1e-2 up_regulated <- res[res$log2FoldChange > 2 & res$padj < 1e-2, ] # Filter down-regulated genes: log2FoldChange < -2 and padj < 1e-2 down_regulated <- res[res$log2FoldChange < -2 & res$padj < 1e-2, ] # Create a new workbook wb <- createWorkbook() # Add the complete dataset as the first sheet addWorksheet(wb, "Complete_Data") writeData(wb, "Complete_Data", res) # Add the up-regulated genes as the second sheet addWorksheet(wb, "Up_Regulated") writeData(wb, "Up_Regulated", up_regulated) # Add the down-regulated genes as the third sheet addWorksheet(wb, "Down_Regulated") writeData(wb, "Down_Regulated", down_regulated) # Save the workbook to a file saveWorkbook(wb, "Gene_Expression_Δsbp_MH_18h_vs_WT_MH_18h.xlsx", overwrite = TRUE) # Set the 'GeneName' column as row.names rownames(res) <- res$GeneName # Drop the 'GeneName' column since it's now the row names res$GeneName <- NULL head(res) #library(EnhancedVolcano) # Assuming res is already sorted and processed png("Δsbp_MH_18h_vs_WT_MH_18h.png", width=1200, height=1200) #max.overlaps = 10 EnhancedVolcano(res, lab = rownames(res), x = 'log2FoldChange', y = 'padj', pCutoff = 1e-2, FCcutoff = 2, title = '', subtitleLabSize = 18, pointSize = 3.0, labSize = 5.0, colAlpha = 1, legendIconSize = 4.0, drawConnectors = TRUE, widthConnectors = 0.5, colConnectors = 'black', subtitle = expression("Δsbp MH 18h versus WT MH 18h")) dev.off() #Annotate the Gene_Expression_xxx_vs_yyy.xlsx in the next steps (see below e.g. Gene_Expression_with_Annotations_Urine_vs_MHB.xlsx) -
Clustering the genes and draw heatmap
#http://xgenes.com/article/article-content/150/draw-venn-diagrams-using-matplotlib/ #http://xgenes.com/article/article-content/276/go-terms-for-s-epidermidis/ # save the Up-regulated and Down-regulated genes into -up.id and -down.id for i in deltasbp_TSB_2h_vs_WT_TSB_2h deltasbp_TSB_4h_vs_WT_TSB_4h deltasbp_TSB_18h_vs_WT_TSB_18h deltasbp_MH_2h_vs_WT_MH_2h deltasbp_MH_4h_vs_WT_MH_4h deltasbp_MH_18h_vs_WT_MH_18h; do echo "cut -d',' -f1-1 ${i}-up.txt > ${i}-up.id"; echo "cut -d',' -f1-1 ${i}-down.txt > ${i}-down.id"; done cat *.id | sort -u > ids #add Gene_Id in the first line, delete the "" #Note that using GeneID as index, rather than GeneName, since .txt contains only GeneID. GOI <- read.csv("ids")$Gene_Id #95 RNASeq.NoCellLine <- assay(rld) #install.packages("gplots") library("gplots") #clustering methods: "ward.D", "ward.D2", "single", "complete", "average" (= UPGMA), "mcquitty" (= WPGMA), "median" (= WPGMC) or "centroid" (= UPGMC). pearson or spearman datamat = RNASeq.NoCellLine[GOI, ] #datamat = RNASeq.NoCellLine write.csv(as.data.frame(datamat), file ="DEGs_heatmap_expression_data.txt") constant_rows <- apply(datamat, 1, function(row) var(row) == 0) if(any(constant_rows)) { cat("Removing", sum(constant_rows), "constant rows.\n") datamat <- datamat[!constant_rows, ] } hr <- hclust(as.dist(1-cor(t(datamat), method="pearson")), method="complete") hc <- hclust(as.dist(1-cor(datamat, method="spearman")), method="complete") mycl = cutree(hr, h=max(hr$height)/1.1) mycol = c("YELLOW", "BLUE", "ORANGE", "MAGENTA", "CYAN", "RED", "GREEN", "MAROON", "LIGHTBLUE", "PINK", "MAGENTA", "LIGHTCYAN", "LIGHTRED", "LIGHTGREEN"); mycol = mycol[as.vector(mycl)] #png("DEGs_heatmap.png", width=900, height=800) #cex.lab=10, labRow="", #png("DEGs_heatmap.png", width=1200, height=1000) #heatmap.2(as.matrix(datamat),Rowv=as.dendrogram(hr),Colv = NA, dendrogram = 'row',labRow="", scale='row',trace='none',col=bluered(75), cexCol=1.8, RowSideColors = mycol, margins=c(10,2), cexRow=1.5, srtCol=34, lhei = c(1, 8), lwid=c(2, 8)) #rownames(datamat) png("DEGs_heatmap.png", width=1200, height=2000) heatmap.2(datamat, Rowv = as.dendrogram(hr), col = bluered(75), scale = "row", RowSideColors = mycol, trace = "none", margin = c(10, 15), # bottom, left sepwidth = c(0, 0), dendrogram = 'row', Colv = 'false', density.info = 'none', labRow = rownames(datamat), cexRow = 1.5, cexCol = 1.5, srtCol = 35, lhei = c(0.2, 4), # reduce top space (was 1 or more) lwid = c(0.4, 4)) # reduce left space (was 1 or more) dev.off() #### cluster members ##### write.csv(names(subset(mycl, mycl == '1')),file='cluster1_YELLOW.txt') write.csv(names(subset(mycl, mycl == '2')),file='cluster2_DARKBLUE.txt') write.csv(names(subset(mycl, mycl == '3')),file='cluster3_DARKORANGE.txt') write.csv(names(subset(mycl, mycl == '4')),file='cluster4_DARKMAGENTA.txt') write.csv(names(subset(mycl, mycl == '5')),file='cluster5_DARKCYAN.txt') #~/Tools/csv2xls-0.4/csv_to_xls.py cluster*.txt -d',' -o DEGs_heatmap_cluster_members.xls #~/Tools/csv2xls-0.4/csv_to_xls.py DEGs_heatmap_expression_data.txt -d',' -o DEGs_heatmap_expression_data.xls; #### (NOT_WORKING) cluster members (adding annotations, note that it does not work for the bacteria, since it is not model-speices and we cannot use mart=ensembl) ##### subset_1<-names(subset(mycl, mycl == '1')) data <- as.data.frame(datamat[rownames(datamat) %in% subset_1, ]) #2575 subset_2<-names(subset(mycl, mycl == '2')) data <- as.data.frame(datamat[rownames(datamat) %in% subset_2, ]) #1855 subset_3<-names(subset(mycl, mycl == '3')) data <- as.data.frame(datamat[rownames(datamat) %in% subset_3, ]) #217 subset_4<-names(subset(mycl, mycl == '4')) data <- as.data.frame(datamat[rownames(datamat) %in% subset_4, ]) # subset_5<-names(subset(mycl, mycl == '5')) data <- as.data.frame(datamat[rownames(datamat) %in% subset_5, ]) # # Initialize an empty data frame for the annotated data annotated_data <- data.frame() # Determine total number of genes total_genes <- length(rownames(data)) # Loop through each gene to annotate for (i in 1:total_genes) { gene <- rownames(data)[i] result <- getBM(attributes = c('ensembl_gene_id', 'external_gene_name', 'gene_biotype', 'entrezgene_id', 'chromosome_name', 'start_position', 'end_position', 'strand', 'description'), filters = 'ensembl_gene_id', values = gene, mart = ensembl) # If multiple rows are returned, take the first one if (nrow(result) > 1) { result <- result[1, ] } # Check if the result is empty if (nrow(result) == 0) { result <- data.frame(ensembl_gene_id = gene, external_gene_name = NA, gene_biotype = NA, entrezgene_id = NA, chromosome_name = NA, start_position = NA, end_position = NA, strand = NA, description = NA) } # Transpose expression values expression_values <- t(data.frame(t(data[gene, ]))) colnames(expression_values) <- colnames(data) # Combine gene information and expression data combined_result <- cbind(result, expression_values) # Append to the final dataframe annotated_data <- rbind(annotated_data, combined_result) # Print progress every 100 genes if (i %% 100 == 0) { cat(sprintf("Processed gene %d out of %d\n", i, total_genes)) } } # Save the annotated data to a new CSV file write.csv(annotated_data, "cluster1_YELLOW.csv", row.names=FALSE) write.csv(annotated_data, "cluster2_DARKBLUE.csv", row.names=FALSE) write.csv(annotated_data, "cluster3_DARKORANGE.csv", row.names=FALSE) write.csv(annotated_data, "cluster4_DARKMAGENTA.csv", row.names=FALSE) write.csv(annotated_data, "cluster5_DARKCYAN.csv", row.names=FALSE) #~/Tools/csv2xls-0.4/csv_to_xls.py cluster*.csv -d',' -o DEGs_heatmap_clusters.xls
KEGG and GO annotations in non-model organisms
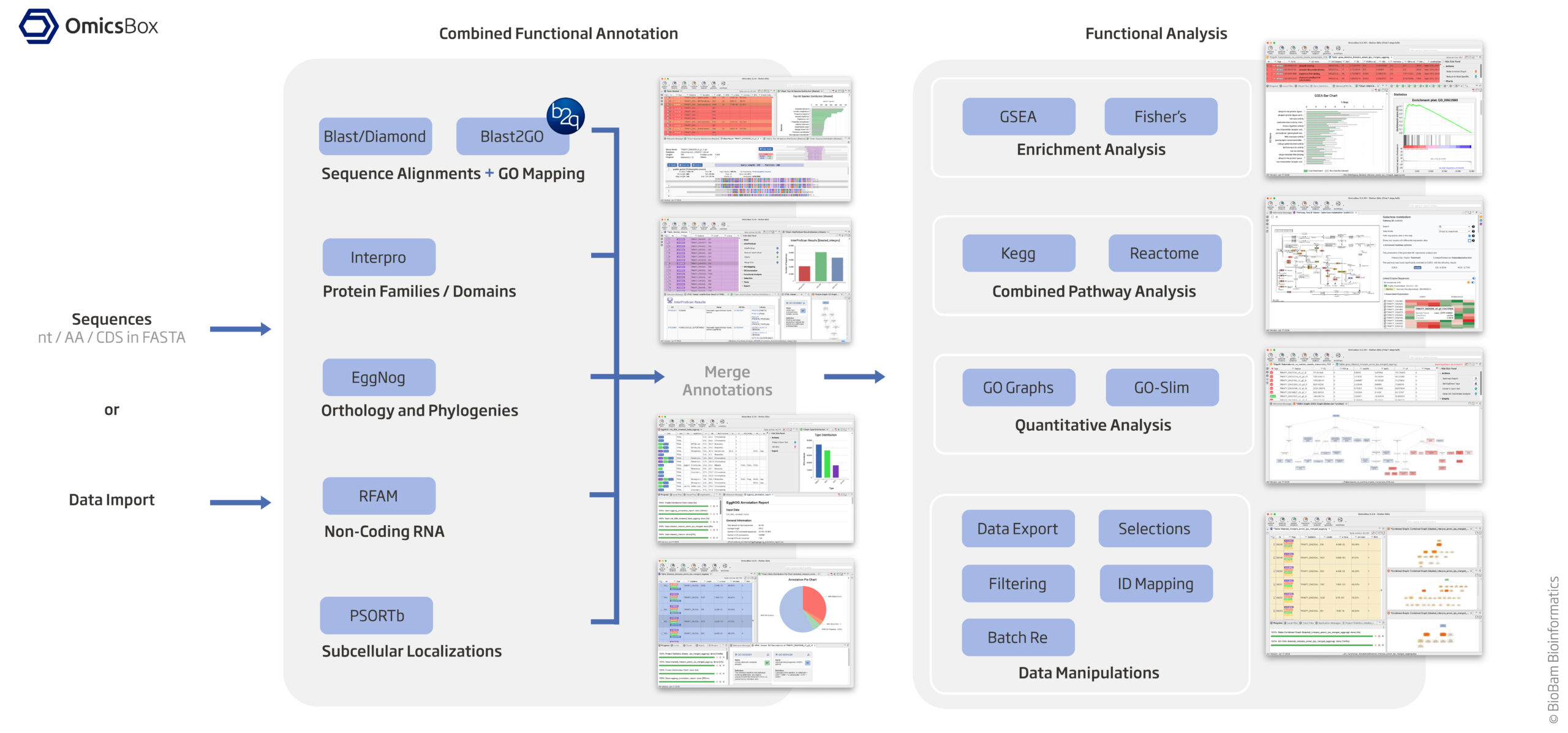
https://www.biobam.com/functional-analysis/
-
Assign KEGG and GO Terms (see diagram above)
Since your organism is non-model, standard R databases (org.Hs.eg.db, etc.) won’t work. You’ll need to manually retrieve KEGG and GO annotations. Option 1 (KEGG Terms): EggNog based on orthology and phylogenies EggNOG-mapper assigns both KEGG Orthology (KO) IDs and GO terms. Install EggNOG-mapper: mamba create -n eggnog_env python=3.8 eggnog-mapper -c conda-forge -c bioconda #eggnog-mapper_2.1.12 mamba activate eggnog_env Run annotation: #diamond makedb --in eggnog6.prots.faa -d eggnog_proteins.dmnd mkdir /home/jhuang/mambaforge/envs/eggnog_env/lib/python3.8/site-packages/data/ download_eggnog_data.py --dbname eggnog.db -y --data_dir /home/jhuang/mambaforge/envs/eggnog_env/lib/python3.8/site-packages/data/ #NOT_WORKING: emapper.py -i CP020463_gene.fasta -o eggnog_dmnd_out --cpu 60 -m diamond[hmmer,mmseqs] --dmnd_db /home/jhuang/REFs/eggnog_data/data/eggnog_proteins.dmnd #Download the protein sequences from Genbank mv ~/Downloads/sequence\ \(3\).txt CP020463_protein_.fasta python ~/Scripts/update_fasta_header.py CP020463_protein_.fasta CP020463_protein.fasta emapper.py -i CP020463_protein.fasta -o eggnog_out --cpu 60 #--resume #----> result annotations.tsv: Contains KEGG, GO, and other functional annotations. #----> 470.IX87_14445: * 470 likely refers to the organism or strain (e.g., Acinetobacter baumannii ATCC 19606 or another related strain). * IX87_14445 would refer to a specific gene or protein within that genome. Extract KEGG KO IDs from annotations.emapper.annotations. Option 2 (GO Terms from 'Blast2GO 5 Basic', saved in blast2go_annot.annot): Using Blast/Diamond + Blast2GO_GUI based on sequence alignment + GO mapping * jhuang@WS-2290C:~/DATA/Data_Michelle_RNAseq_2025$ ~/Tools/Blast2GO/Blast2GO_Launcher setting the workspace "mkdir ~/b2gWorkspace_Michelle_RNAseq_2025"; cp /mnt/md1/DATA/Data_Michelle_RNAseq_2025/results/star_salmon/degenes/CP020463_protein.fasta ~/b2gWorkspace_Michelle_RNAseq_2025 * 'Load protein sequences' (Tags: NONE, generated columns: Nr, SeqName) by choosing the file CP020463_protein.fasta as input --> * Buttons 'blast' at the NCBI (Parameters: blastp, nr, ...) (Tags: BLASTED, generated columns: Description, Length, #Hits, e-Value, sim mean), QBlast finished with warnings! Blasted Sequences: 2084 Sequences without results: 105 Check the Job log for details and try to submit again. Restarting QBlast may result in additional results, depending on the error type. "Blast (CP020463_protein) Done" * Button 'mapping' (Tags: MAPPED, generated columns: #GO, GO IDs, GO Names), "Mapping finished - Please proceed now to annotation." "Mapping (CP020463_protein) Done" "Mapping finished - Please proceed now to annotation." * Button 'annot' (Tags: ANNOTATED, generated columns: Enzyme Codes, Enzyme Names), "Annotation finished." * Used parameter 'Annotation CutOff': The Blast2GO Annotation Rule seeks to find the most specific GO annotations with a certain level of reliability. An annotation score is calculated for each candidate GO which is composed by the sequence similarity of the Blast Hit, the evidence code of the source GO and the position of the particular GO in the Gene Ontology hierarchy. This annotation score cutoff select the most specific GO term for a given GO branch which lies above this value. * Used parameter 'GO Weight' is a value which is added to Annotation Score of a more general/abstract Gene Ontology term for each of its more specific, original source GO terms. In this case, more general GO terms which summarise many original source terms (those ones directly associated to the Blast Hits) will have a higher Annotation Score. "Annotation (CP020463_protein) Done" "Annotation finished." or blast2go_cli_v1.5.1 (NOT_USED) #https://help.biobam.com/space/BCD/2250407989/Installation #see ~/Scripts/blast2go_pipeline.sh Option 3 (GO Terms from 'Blast2GO 5 Basic', saved in blast2go_annot.annot2): Interpro based protein families / domains --> Button interpro * Button 'interpro' (Tags: INTERPRO, generated columns: InterPro IDs, InterPro GO IDs, InterPro GO Names) --> "InterProScan Finished - You can now merge the obtained GO Annotations." "InterProScan (CP020463_protein) Done" "InterProScan Finished - You can now merge the obtained GO Annotations." MERGE the results of InterPro GO IDs (Option 3) to GO IDs (Option 2) and generate final GO IDs * Button 'interpro'/'Merge InterProScan GOs to Annotation' --> "Merge (add and validate) all GO terms retrieved via InterProScan to the already existing GO annotation." "Merge InterProScan GOs to Annotation (CP020463_protein) Done" "Finished merging GO terms from InterPro with annotations." "Maybe you want to run ANNEX (Annotation Augmentation)." #* Button 'annot'/'ANNEX' --> "ANNEX finished. Maybe you want to do the next step: Enzyme Code Mapping." File -> Export -> Export Annotations -> Export Annotations (.annot, custom, etc.) #~/b2gWorkspace_Michelle_RNAseq_2025/blast2go_annot.annot is generated! #-- before merging (blast2go_annot.annot) -- #H0N29_18790 GO:0004842 ankyrin repeat domain-containing protein #H0N29_18790 GO:0085020 #-- after merging (blast2go_annot.annot2) --> #H0N29_18790 GO:0031436 ankyrin repeat domain-containing protein #H0N29_18790 GO:0070531 #H0N29_18790 GO:0004842 #H0N29_18790 GO:0005515 #H0N29_18790 GO:0085020 cp blast2go_annot.annot blast2go_annot.annot2 Option 4 (NOT_USED): RFAM for non-colding RNA Option 5 (NOT_USED): PSORTb for subcellular localizations Option 6 (NOT_USED): KAAS (KEGG Automatic Annotation Server) * Go to KAAS * Upload your FASTA file. * Select an appropriate gene set. * Download the KO assignments. -
Find the Closest KEGG Organism Code (NOT_USED)
Since your species isn't directly in KEGG, use a closely related organism. * Check available KEGG organisms: library(clusterProfiler) library(KEGGREST) kegg_organisms <- keggList("organism") Pick the closest relative (e.g., zebrafish "dre" for fish, Arabidopsis "ath" for plants). # Search for Acinetobacter in the list grep("Acinetobacter", kegg_organisms, ignore.case = TRUE, value = TRUE) # Gammaproteobacteria #Extract KO IDs from the eggnog results for "Acinetobacter baumannii strain ATCC 19606" -
Find the Closest KEGG Organism for a Non-Model Species (NOT_USED)
If your organism is not in KEGG, search for the closest relative: grep("fish", kegg_organisms, ignore.case = TRUE, value = TRUE) # Example search For KEGG pathway enrichment in non-model species, use "ko" instead of a species code (the code has been intergrated in the point 4): kegg_enrich <- enrichKEGG(gene = gene_list, organism = "ko") # "ko" = KEGG Orthology -
Perform KEGG and GO Enrichment in R (under dir ~/DATA/Data_Tam_RNAseq_2025_LB_vs_Mac_ATCC19606/results/star_salmon/degenes)
#BiocManager::install("GO.db") #BiocManager::install("AnnotationDbi") # Load required libraries library(openxlsx) # For Excel file handling library(dplyr) # For data manipulation library(tidyr) library(stringr) library(clusterProfiler) # For KEGG and GO enrichment analysis #library(org.Hs.eg.db) # Replace with appropriate organism database library(GO.db) library(AnnotationDbi) setwd("~/DATA/Data_Michelle_RNAseq_2025/results/star_salmon/degenes") # PREPARING go_terms and ec_terms: annot_* file: cut -f1-2 -d$'\t' blast2go_annot.annot2 > blast2go_annot.annot2_ # PREPARING eggnog_out.emapper.annotations.txt from eggnog_out.emapper.annotations by removing ## lines and renaming #query to query #(plot-numpy1) jhuang@WS-2290C:~/DATA/Data_Tam_RNAseq_2024_AUM_MHB_Urine_ATCC19606$ diff eggnog_out.emapper.annotations eggnog_out.emapper.annotations.txt #1,5c1 #< ## Thu Jan 30 16:34:52 2025 #< ## emapper-2.1.12 #< ## /home/jhuang/mambaforge/envs/eggnog_env/bin/emapper.py -i CP059040_protein.fasta -o eggnog_out --cpu 60 #< ## #< #query seed_ortholog evalue score eggNOG_OGs max_annot_lvl COG_category Description Preferred_name GOs EC KEGG_ko KEGG_Pathway KEGG_Module KEGG_Reaction KEGG_rclass BRITE KEGG_TC CAZy BiGG_Reaction PFAMs #--- #> query seed_ortholog evalue score eggNOG_OGs max_annot_lvl COG_category Description Preferred_name GOs EC KEGG_ko KEGG_Pathway KEGG_Module KEGG_Reaction KEGG_rclass BRITE KEGG_TC CAZy BiGG_Reaction PFAMs #3620,3622d3615 #< ## 3614 queries scanned #< ## Total time (seconds): 8.176708459854126 # Step 1: Load the blast2go annotation file with a check for missing columns annot_df <- read.table("/home/jhuang/b2gWorkspace_Michelle_RNAseq_2025/blast2go_annot.annot2_", header = FALSE, sep = "\t", stringsAsFactors = FALSE, fill = TRUE) # If the structure is inconsistent, we can make sure there are exactly 3 columns: colnames(annot_df) <- c("GeneID", "Term") # Step 2: Filter and aggregate GO and EC terms as before go_terms <- annot_df %>% filter(grepl("^GO:", Term)) %>% group_by(GeneID) %>% summarize(GOs = paste(Term, collapse = ","), .groups = "drop") ec_terms <- annot_df %>% filter(grepl("^EC:", Term)) %>% group_by(GeneID) %>% summarize(EC = paste(Term, collapse = ","), .groups = "drop") # Key Improvements: # * Looped processing of all 6 input files to avoid redundancy. # * Robust handling of empty KEGG and GO enrichment results to prevent contamination of results between iterations. # * File-safe output: Each dataset creates a separate Excel workbook with enriched sheets only if data exists. # * Error handling for GO term descriptions via tryCatch. # * Improved clarity and modular structure for easier maintenance and future additions. # Define the filenames and output suffixes file_list <- c( "deltasbp_TSB_2h_vs_WT_TSB_2h-all.csv", "deltasbp_TSB_4h_vs_WT_TSB_4h-all.csv", "deltasbp_TSB_18h_vs_WT_TSB_18h-all.csv", "deltasbp_MH_2h_vs_WT_MH_2h-all.csv", "deltasbp_MH_4h_vs_WT_MH_4h-all.csv", "deltasbp_MH_18h_vs_WT_MH_18h-all.csv" ) file_name = "deltasbp_MH_18h_vs_WT_MH_18h-all.csv" #for (file_name in file_list) { message("Processing file: ", file_name) # Clear variables from previous iteration res <- NULL res_updated <- NULL up_regulated <- NULL down_regulated <- NULL gene_list_kegg_up <- NULL gene_list_kegg_down <- NULL kegg_enrichment_up <- NULL kegg_enrichment_down <- NULL kegg_enrichment_up_df <- NULL kegg_enrichment_down_df <- NULL gene_list_go_up <- NULL go_terms_df_up <- NULL gene_list_go_down <- NULL go_terms_df_down <- NULL background_genes <- NULL go_annotation <- NULL go_enrichment_up <- NULL go_enrichment_down <- NULL wb <- NULL # Now load and process fresh for this file res <- read.csv(file_name) # Replace empty GeneName with modified GeneID res$GeneName <- ifelse( res$GeneName == "" | is.na(res$GeneName), gsub("gene-", "", res$GeneID), res$GeneName ) # Remove duplicates by smallest padj res <- res %>% group_by(GeneName) %>% slice_min(padj, with_ties = FALSE) %>% ungroup() %>% as.data.frame() # Sort by padj ascending, then log2FoldChange descending res <- res[order(res$padj, -res$log2FoldChange), ] # Load eggnog data eggnog_data <- read.delim("~/DATA/Data_Michelle_RNAseq_2025/eggnog_out.emapper.annotations.txt", header = TRUE, sep = "\t") res$GeneID <- gsub("gene-", "", res$GeneID) res <- res %>% left_join(eggnog_data, by = c("GeneID" = "query")) # Join with go_terms and ec_terms (assumed preloaded) res_updated <- res %>% left_join(go_terms, by = "GeneID") %>% left_join(ec_terms, by = "GeneID") %>% select(-EC.x, -GOs.x) %>% rename(EC = EC.y, GOs = GOs.y) # Filter up/down regulated genes up_regulated <- res_updated %>% filter(log2FoldChange > 2, padj < 0.01) down_regulated <- res_updated %>% filter(log2FoldChange < -2, padj < 0.01) # Prepare KEGG enrichment for up-regulated gene_list_kegg_up <- gsub("ko:", "", up_regulated$KEGG_ko) gene_list_kegg_up <- gene_list_kegg_up[!is.na(gene_list_kegg_up) & gene_list_kegg_up != ""] kegg_enrichment_up <- enrichKEGG(gene = gene_list_kegg_up, organism = 'ko') # Prepare KEGG enrichment for down-regulated gene_list_kegg_down <- gsub("ko:", "", down_regulated$KEGG_ko) gene_list_kegg_down <- gene_list_kegg_down[!is.na(gene_list_kegg_down) & gene_list_kegg_down != ""] kegg_enrichment_down <- enrichKEGG(gene = gene_list_kegg_down, organism = 'ko') # Function to get data.frame from enrichKEGG or return empty data.frame if no results get_enrich_df <- function(enrich_obj) { if (is.null(enrich_obj) || length(enrich_obj) == 0) { return(data.frame()) } res_df <- enrich_obj@result if (nrow(res_df) == 0) return(data.frame()) return(res_df) } kegg_enrichment_up_df <- get_enrich_df(kegg_enrichment_up) kegg_enrichment_down_df <- get_enrich_df(kegg_enrichment_down) #COPY_BREAK_POINT # Prepare GO enrichment gene_list_go_up <- up_regulated$GeneID gene_list_go_down <- down_regulated$GeneID background_genes <- res_updated$GeneID go_annotation <- res_updated[, c("GOs", "GeneID")] %>% separate_rows(GOs, sep = ",") go_enrichment_up <- enricher( gene = gene_list_go_up, TERM2GENE = go_annotation, pvalueCutoff = 0.05, pAdjustMethod = "BH", universe = background_genes ) %>% as.data.frame() go_enrichment_down <- enricher( gene = gene_list_go_down, TERM2GENE = go_annotation, pvalueCutoff = 0.05, pAdjustMethod = "BH", universe = background_genes ) %>% as.data.frame() # Check if enrichment results are empty and print messages if (nrow(kegg_enrichment_up_df) == 0) { message("No KEGG enrichment results for UP genes in file: ", file_name) } else { message("KEGG enrichment UP results (head) for file: ", file_name) print(head(kegg_enrichment_up_df)) } if (nrow(kegg_enrichment_down_df) == 0) { message("No KEGG enrichment results for DOWN genes in file: ", file_name) } else { message("KEGG enrichment DOWN results (head) for file: ", file_name) print(head(kegg_enrichment_down_df)) } if (nrow(go_enrichment_up) == 0) { message("No GO enrichment results for UP genes in file: ", file_name) } else { # Create a lookup table: GOID -> Term go_terms_df_up <- AnnotationDbi::select(GO.db, keys = go_enrichment_up$ID, columns = c("TERM"), keytype = "GOID") # Join back the term descriptions to your enrichment results go_enrichment_up <- go_enrichment_up %>% left_join(go_terms_df_up, by = c("ID" = "GOID")) %>% # Replace Description column with TERM from GO.db mutate(Description = TERM) %>% select(-TERM) message("GO enrichment UP results for file (head): ", file_name) print(head(go_enrichment_up)) } if (nrow(go_enrichment_down) == 0) { message("No GO enrichment results for DOWN genes in file: ", file_name) } else { # Create a lookup table: GOID -> Term go_terms_df_down <- AnnotationDbi::select(GO.db, keys = go_enrichment_down$ID, columns = c("TERM"), keytype = "GOID") # Join back the term descriptions to your enrichment results go_enrichment_down <- go_enrichment_down %>% left_join(go_terms_df_down, by = c("ID" = "GOID")) %>% # Replace Description column with TERM from GO.db mutate(Description = TERM) %>% select(-TERM) message("GO enrichment DOWN results for file (head): ", file_name) print(head(go_enrichment_down)) } # Save results to workbook wb <- createWorkbook() addWorksheet(wb, "Complete_Data") writeData(wb, "Complete_Data", res_updated) addWorksheet(wb, "Up_Regulated") writeData(wb, "Up_Regulated", up_regulated) addWorksheet(wb, "Down_Regulated") writeData(wb, "Down_Regulated", down_regulated) addWorksheet(wb, "KEGG_Enrichment_Up") writeData(wb, "KEGG_Enrichment_Up", kegg_enrichment_up_df) addWorksheet(wb, "KEGG_Enrichment_Down") writeData(wb, "KEGG_Enrichment_Down", kegg_enrichment_down_df) addWorksheet(wb, "GO_Enrichment_Up") writeData(wb, "GO_Enrichment_Up", go_enrichment_up) addWorksheet(wb, "GO_Enrichment_Down") writeData(wb, "GO_Enrichment_Down", go_enrichment_down) output_file <- paste0("DEG_KEGG_GO_", gsub("-all.csv", "", file_name), ".xlsx") saveWorkbook(wb, output_file, overwrite = TRUE) message("Finished processing: ", file_name, "\n\n") #}
点赞本文的读者
还没有人对此文章表态
本文有评论
没有评论
看文章,发评论,不要沉默
最受欢迎文章
- Motif Discovery in Biological Sequences: A Comparison of MEME and HOMER
- Why Do Significant Gene Lists Change After Adding Additional Conditions in Differential Gene Expression Analysis?
- Calling peaks using findPeaks of HOMER
- PiCRUST2 Pipeline for Functional Prediction and Pathway Analysis in Metagenomics
- Updating Human Gene Identifiers using Ensembl BioMart: A Step-by-Step Guide
- Should the inputs for GSVA be normalized or raw?
- pheatmap vs heatmap.2
- Kraken2 Installation and Usage Guide
- Setup conda environments
- File format for single channel analysis of Agilent microarray data with Limma?
最新文章
- Processing Data_Michelle_RNAseq_2025
- 从共生到致病:金黄色葡萄球菌在鼻腔定植与人工关节感染过程中的转录表达谱
- 外排泵缺失揭示鲍曼不动杆菌 ATCC19606 对氯霉素的菌株特异性转录组适应机制
- 亲子鉴定中的STR分析简要概述
最多评论文章
- Updating Human Gene Identifiers using Ensembl BioMart: A Step-by-Step Guide
- The top 10 genes
- Retrieving KEGG Genes Using Bioservices in Python
推荐相似文章
Enhanced Visualization of Gene Presence for the Selected Genes in Bongarts_S.epidermidis_HDRNA
RNA-seq Tam on CP059040.1 (Acinetobacter baumannii strain ATCC 19606)
Variant Calling for Herpes Simplex Virus 1 from Patient Sample Using Capture Probe Sequencing